 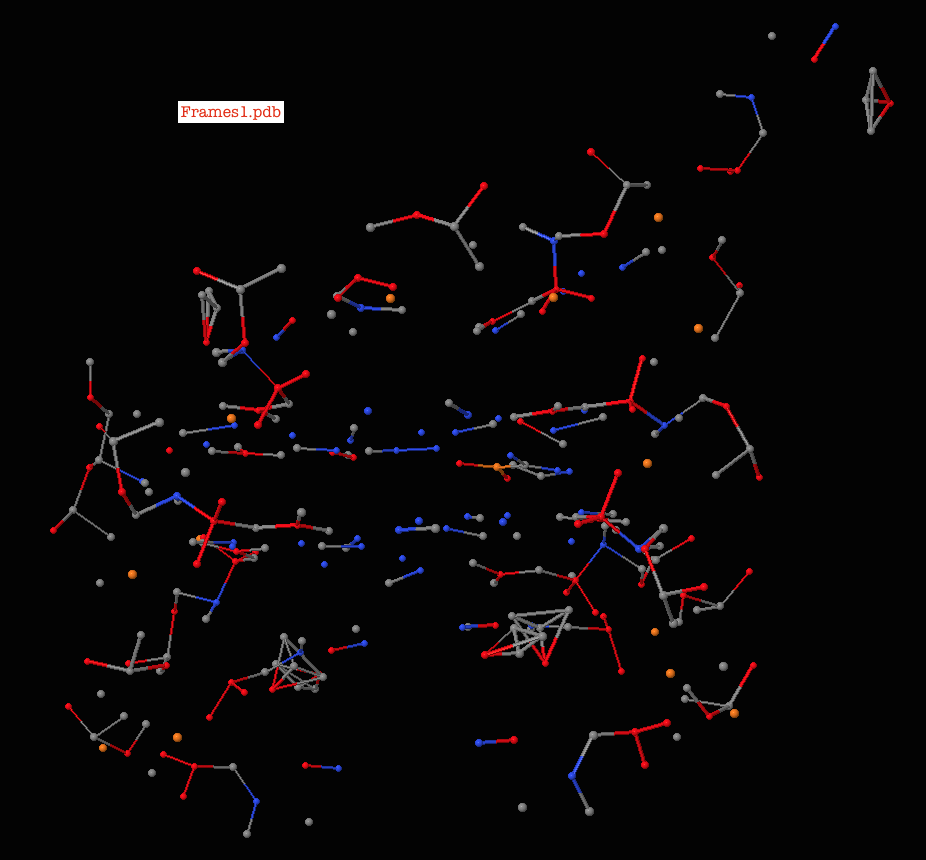
DALTON: The molden file from dalton cannot be opened by jmol."o2_state_tpssd3_prod_oh_opt_cosmo.xyz" "o2_state_tpssd3_o_opt_cosmo.xyz"Įcho "load structure.xyz spacefill 0.2" > orbital_$i".spt"Įcho 'isosurface sign "'$i.cube'" translucent 0.4' > orbital_$i".spt"Įcho "write jpg $i.jpg" > orbital_$i".spt" "o2_state_tpssd3_react_oo_opt_cosmo.xyz" "o2_state_tpssd3_oh2_opt_cosmo.xyz" "o2_state_tpssd3_react_oh_opt_cosmo_singlet.xyz" Next make the representations of the bases and exclude those that are in the background of the final image. Load FILES "o2_state_tpssd3_ooh2_opt_cosmo.xyz" The following java script can be inserted in the Jmol scripting window - It loads the files rotate them (adjust this according to your needs), writes the distances between atoms 49/50, 49/1, 49/10 and 49/30 and prints a frame in jpg format.Script to generate many MOs for i in do cubegen 0 MO="$i" orbitals.fchk "$i".cube done Jmol structures and molecular orbitals Note for an unrestricted calculation, use "AMO" or "BMO" instead. Use the utility program cubegen cubegen 0 MO= gaussian.fchk.Note: make sure the gaussian module is loaded (aurora: module load gaussian/16.B.01-AVX2) Transform the ".chk" with the utility program formchk gaussian.chk gaussian.fchk. New in vmd-python > 3.0.0, you can access these attributes directly from the atomsel object to both get and set For example: protein atomsel('protein') protein.chain 'A'.Run a gaussian wave function optimization and make sure you obtain the ".chk" file, e.g.We focus here on generation of cube files that can be visualized with Jmol (see below). In gaussian, the can be generated in different ways and visualized with Jmol (see below). They are particularly useful for interpreting electronic spectroscopy (i.e. Molecular orbitals are often useful for understanding the electronic structure of a molecule. xyz or pdb file can be written out with from File => Save Coordinates.IMPORTANT: The align will happend according to the "top" molecule (marked with "T" in the VMD main window - this molecule will have unchanged coordinates) It only works if there are equally many of the atom types in the fwo files!). (note that they should have the same name in the pdb/xyz files that are compared. 2) Select the atom of interest by clicking on it with left mouse button. Now type for the atoms that should be aligned (in box upper left) name ATOM1 ATOM2 ATOM3 etc. 1) Press key '1' to enter atom picking mode.Check that it is the right molecules that are read in (try "Add all"). Extensions => Analysis => RMSD Trajectory Tool.Note: this can also be done (often faster) with Maestro "Quick Align" However, if a selection is performed by index, as with index 1, VMD will select the second atom, and the. 
Type vmd > set sel4 įor all examples, write the pdb using writepdb selection: vmd > $selX writepdb pdb_out.pdb VMD uses 0-based indexing and VMDselect adjusts that.Select whole molecules, here waters - and only waters Save pdb with selection of atoms within 5 Å of another atom, here "Mn" type command set sel1 Ĭheck that the number of atoms fit by command $sel1numĮxample 2.Saving pdbs with different selections (examples) within 5 of name FE (selects all atoms withn 5 Å of FE).name CA (typically selects alpha-C in protiens if the "normal" pdb name for alpha carbons are used).Graphics => Representations: The menu "Selected Atoms" can be used. Vmd atom selection serial#the PDB would look like serial 1 to 100 You. Selected Atoms box in the Graphical Representations window (Graphics Representations). Read in pdb or xyz structure: vmd pdb_name.pdb or vmd xyz_name.xyzĪfter starting a VMD shell (see above) go to There are two numbering schemes for atoms in VMD, one starts numbering at 1 (serial) and the other at 0 (index).VMD can be found here: starting vmd and loading structures and Schulten, K., "VMD - Visual Molecular Dynamics", J. VMD development is supported by the National Institutes of Health grant numbers NIH 9P41GM104601 and 5R01GM098243-02. 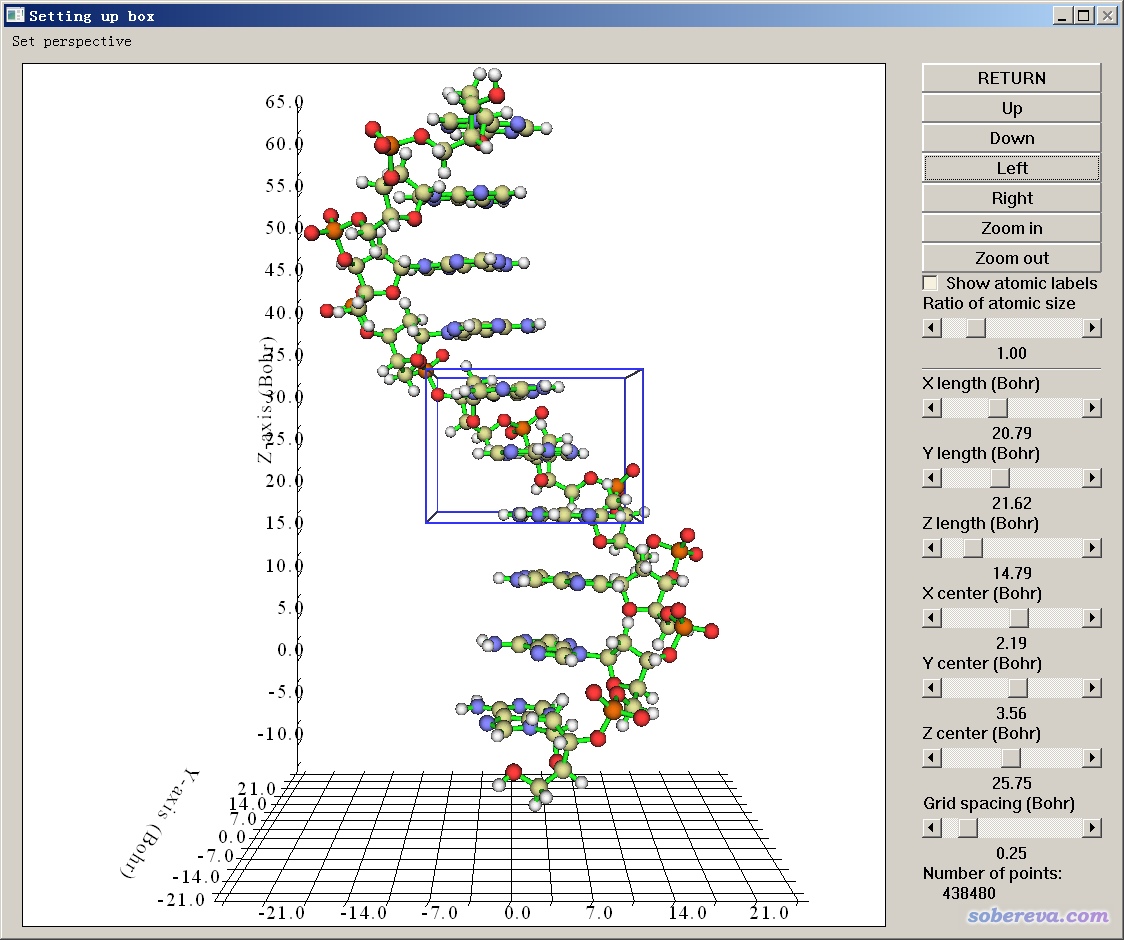
Vmd atom selection how to#Useful commands and features of the powerfull VMD software (Humphrey, W., Dalke, A. This guide documents the user interfaces displaying and grapically manipulating molecules, and describes how to use the scripting interfaces for analysis and to customize the behavior of VMD. Visualization tools for quantum chemistry Up: VMD User's Guide Previous: Future Plans Contents 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
